The example is made of a single source file (one version in each supported programming language), this file loads its configuration and PDI specification tree from a YAML file specified on the command line. Just by replacing the configuration file, the example program will use different PDI plugins, different libraries and different I/O operations.
The example implements implements a simple Heat equation solver using an explicit forward finite difference scheme parallelized with MPI in 2 dimensions.
The data is stored in a 2D array in which each point represents the temperature. In every iteration, the value at the next iteration of each cell is computed by applying a cross stencil using the values of the cell and of its neighbours (top, bottom, left and right) at the current iteration:
In order not to override the cells while processing we use two arrays to store the values, one for the current iteration (cur in the code) and one for the next iteration (next in the code). These computations are written in the iter function.
This code is parallelized with MPI . Let's split that matrix by MPI processes. Each process will compute part of global matrix.
For example: matrix 16 x 16 integers and 16 MPI processes gives submatrices of 4 x 4 integers for every process. We have to add to our global matrix one row above and below, column to the left and right to be able to compute border cells. In our example row on the top has some value (”x”) bigger than 0 (representing source of heat):
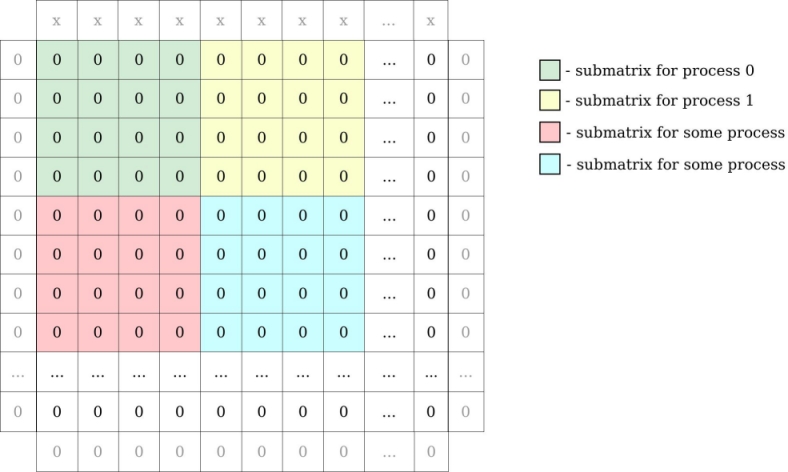
MPI processes need to exchange information about their local matrix border cells (communicate with neighbours to exchange row/column of matrix). Each MPI process will have a local matrix:
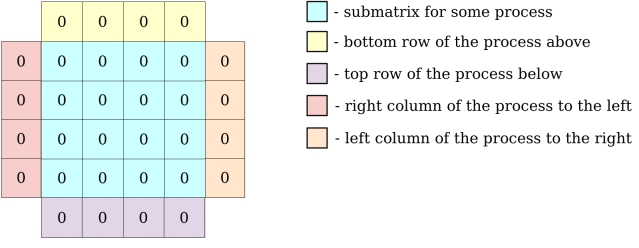
All the communications instructions are written in exchange function.
Now, when we know the algorithm, we can focus on analysing decl_hdf5.yaml specification tree. Fisrt 3 maps defined will not be seen to PDI:
duration is the value in seconds how long the application will run.datasize is size of our global matrix.parallelism defines the number of MPI processes in each dimension.Next, we have defined data and metadata:
In source file we will extract the pdi map and pass it as PDI_init argument.
iter will hold the current iteration number.dsize will hold the size of local matrix of each MPI process.psize will hold number of processes in dimensions.pcoord will hold coordinates for each process.main_filed is the local matrix for each process.Let's take a closer look at C source code.
As mentioned before, we extract the pdi subtree and pass it to PDI_init.
We did not defined mpi_comm data in yaml, so this line will have no effect:
The same goes for all PDI calls with data we didn't defined.
Here we are reading global matrix size from specification tree. Similar with parallelism and duration.
After calculating the local matrix sizes and coordinates, we expose them:
At the beginning of each iteration, we call multiexpose:
Above instruction will share iter and main_field, call newiter event and then reclaim main_field and iter. This is the place when plugins will read/write our data.
We have covered the logic behind the PDI example. Now you can start the Hands-on tutorial.